| Bos taurus Gene: BT.28253 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Summary | |||||||||||||
| InnateDB Gene | IDBG-637232.3 | ||||||||||||
| Last Modified | 2014-10-13 [Report errors or provide feedback] | ||||||||||||
| Gene Symbol | BT.28253 | ||||||||||||
| Gene Name | gamma-glutamyltranspeptidase 1 | ||||||||||||
| Synonyms | |||||||||||||
| Species | Bos taurus | ||||||||||||
| Ensembl Gene | ENSBTAG00000046900 | ||||||||||||
| Encoded Proteins | |||||||||||||
| Protein Structure | |||||||||||||
| Useful resources | Stemformatics EHFPI ImmGen | ||||||||||||
| Entrez Gene | |||||||||||||
| Summary |
This gene does not have any Entrez summary - the following is the summary from its human ortholog ENSG00000100031:
GGT2 belongs to the gamma-glutamyltransferase (GGT; EC 2.3.2.2) gene family. GGT is a membrane-bound extracellular enzyme that cleaves gamma-glutamyl peptide bonds in glutathione and other peptides and transfers the gamma-glutamyl moiety to acceptors. GGT is also key to glutathione homeostasis because it provides substrates for glutathione synthesis (Heisterkamp et al., 2008 [PubMed 18357469]).[supplied by OMIM, Oct 2008] The enzyme encoded by this gene catalyzes the transfer of the glutamyl moiety of glutathione to a variety of amino acids and dipeptide acceptors. The enzyme is composed of a heavy chain and a light chain, which are derived from a single precursor protein, and is present in tissues involved in absorption and secretion. This enzyme is a member of the gamma-glutamyltransferase protein family, of which many members have not yet been fully characterized and some of which may represent pseudogenes. This gene is classified as type I gamma-glutamyltransferase. Multiple alternatively spliced variants, encoding the same protein, have been identified. [provided by RefSeq, Oct 2008] The enzyme encoded by this gene is a type I gamma-glutamyltransferase that catalyzes the transfer of the glutamyl moiety of glutathione to a variety of amino acids and dipeptide acceptors. The enzyme is composed of a heavy chain and a light chain, which are derived from a single precursor protein. It is expressed in tissues involved in absorption and secretion and may contribute to the etiology of diabetes and other metabolic disorders. Multiple alternatively spliced variants have been identified. There are a number of related genes present on chromosomes 20 and 22, and putative pseudogenes for this gene on chromosomes 2, 13, and 22. [provided by RefSeq, Jan 2014] |
||||||||||||
| Gene Information | |||||||||||||
| Type | Protein coding | ||||||||||||
| Genomic Location | Chromosome 17:73437809-73452741 | ||||||||||||
| Strand | Reverse strand | ||||||||||||
| Band | |||||||||||||
| Transcripts |
|
||||||||||||
| Interactions | |||||||||||||
| Number of Interactions |
This gene and/or its encoded proteins are associated with 0 experimentally validated interaction(s) in this database.
They are also associated with 2 interaction(s) predicted by orthology.
|
||||||||||||
| Gene Ontology | |||||||||||||
Molecular Function |
|
||||||||||||
| Biological Process |
|
||||||||||||
| Cellular Component |
|
||||||||||||
| Orthologs | |||||||||||||
|
Species
Homo sapiens
Mus musculus
|
Gene ID
Gene Order
|
||||||||||||
| Pathway Predictions based on Human Orthology Data | |||||||||||||
| NETPATH | |||||||||||||
| REACTOME |
Synthesis of Leukotrienes (LT) and Eoxins (EX) pathway
Arachidonic acid metabolism pathway
Glutathione synthesis and recycling pathway
Glutathione conjugation pathway
Metabolism of lipids and lipoproteins pathway
Aflatoxin activation and detoxification pathway
Phase II conjugation pathway
Metabolism pathway
Biological oxidations pathway
Aflatoxin activation and detoxification pathway
Phase II conjugation pathway
Metabolism pathway
Metabolism of lipids and lipoproteins pathway
Glutathione conjugation pathway
Glutathione synthesis and recycling pathway
Biological oxidations pathway
Arachidonic acid metabolism pathway
Synthesis of Leukotrienes (LT) and Eoxins (EX) pathway
|
||||||||||||
| KEGG |
Arachidonic acid metabolism pathway
Cyanoamino acid metabolism pathway
Taurine and hypotaurine metabolism pathway
Glutathione metabolism pathway
Glutathione metabolism pathway
Arachidonic acid metabolism pathway
Taurine and hypotaurine metabolism pathway
Cyanoamino acid metabolism pathway
|
||||||||||||
| INOH |
Prostaglandin Leukotriene metabolism pathway
|
||||||||||||
| PID NCI | |||||||||||||
| Cross-References | |||||||||||||
| SwissProt | |||||||||||||
| TrEMBL | |||||||||||||
| UniProt Splice Variant | |||||||||||||
| Entrez Gene | |||||||||||||
| UniGene | Bt.28253 | ||||||||||||
| RefSeq | NM_001206214 XM_005218251 XM_005218252 XM_005218253 XM_005218254 | ||||||||||||
| HUGO | |||||||||||||
| OMIM | |||||||||||||
| CCDS | |||||||||||||
| HPRD | |||||||||||||
| IMGT | |||||||||||||
| EMBL | |||||||||||||
| GenPept | |||||||||||||
| RNA Seq Atlas | |||||||||||||
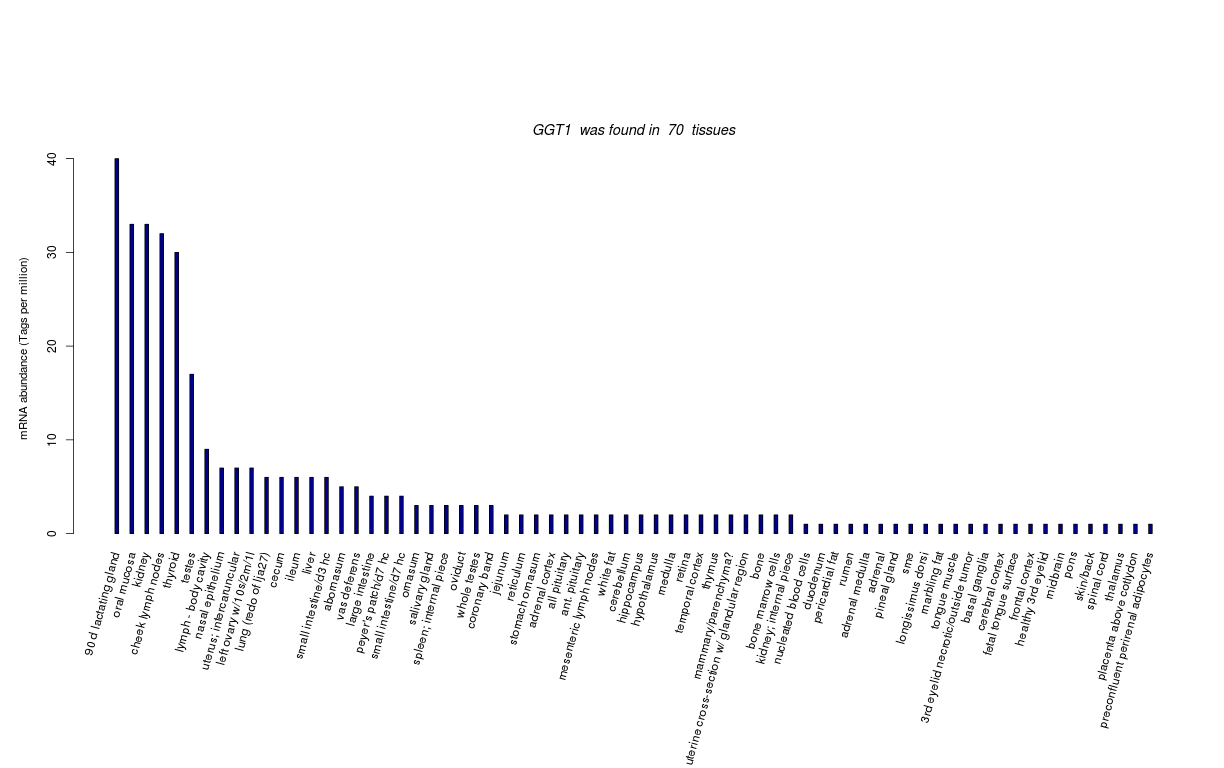
| Transcript Frequencies | |||||||||||||
| Tag Count based mRNA-Abundances across 87 different Tissues (TPM).
Based on Data from Bovine Gene Atlas |
(Move your mouse over the image to view a more detailed version) |
||||||||||||