| Bos taurus Gene: BT.84824 | |||||||
|---|---|---|---|---|---|---|---|
| Summary | |||||||
| InnateDB Gene | IDBG-645684.3 | ||||||
| Last Modified | 2014-10-13 [Report errors or provide feedback] | ||||||
| Gene Symbol | BT.84824 | ||||||
| Gene Name | calcium-dependent phospholipase A2 precursor | ||||||
| Synonyms | |||||||
| Species | Bos taurus | ||||||
| Ensembl Gene | ENSBTAG00000039122 | ||||||
| Encoded Proteins |
Uncharacterized protein
|
||||||
| Protein Structure | |||||||
| Useful resources | Stemformatics EHFPI ImmGen | ||||||
| Entrez Gene | |||||||
| Summary |
This gene does not have any Entrez summary - the following is the summary from its human ortholog ENSG00000127472:
This gene is a member of the secretory phospholipase A2 family. It is located in a tightly-linked cluster of secretory phospholipase A2 genes on chromosome 1. The encoded enzyme catalyzes the hydrolysis of membrane phospholipids to generate lysophospholipids and free fatty acids including arachidonic acid. It preferentially hydrolyzes linoleoyl-containing phosphatidylcholine substrates. Secretion of this enzyme is thought to induce inflammatory responses in neighboring cells. Alternatively spliced transcript variants have been found, but their full-length nature has not been determined. [provided by RefSeq, Jul 2008] |
||||||
| Gene Information | |||||||
| Type | Protein coding | ||||||
| Genomic Location | Chromosome 2:133224531-133246491 | ||||||
| Strand | Reverse strand | ||||||
| Band | |||||||
| Transcripts |
|
||||||
| Interactions | |||||||
| Number of Interactions |
This gene and/or its encoded proteins are associated with 0 experimentally validated interaction(s) in this database.
|
||||||
| Gene Ontology | |||||||
Molecular Function |
|
||||||
| Biological Process |
|
||||||
| Cellular Component |
|
||||||
| Orthologs | |||||||
|
Species
Homo sapiens
Mus musculus
|
Gene ID
Gene Order
|
||||||
| Pathway Predictions based on Human Orthology Data | |||||||
| NETPATH | |||||||
| REACTOME |
Acyl chain remodelling of PC pathway
Acyl chain remodelling of PI pathway
Acyl chain remodelling of PS pathway
Synthesis of PA pathway
Acyl chain remodelling of PG pathway
Acyl chain remodelling of PE pathway
Metabolism of lipids and lipoproteins pathway
Glycerophospholipid biosynthesis pathway
Phospholipid metabolism pathway
Metabolism pathway
|
||||||
| KEGG |
alpha-Linolenic acid metabolism pathway
Arachidonic acid metabolism pathway
Glycerophospholipid metabolism pathway
Linoleic acid metabolism pathway
Ether lipid metabolism pathway
Vascular smooth muscle contraction pathway
Fat digestion and absorption pathway
Pancreatic secretion pathway
Glycerophospholipid metabolism pathway
Arachidonic acid metabolism pathway
Ether lipid metabolism pathway
Linoleic acid metabolism pathway
alpha-Linolenic acid metabolism pathway
Vascular smooth muscle contraction pathway
Pancreatic secretion pathway
Fat digestion and absorption pathway
|
||||||
| INOH | |||||||
| PID NCI | |||||||
| Cross-References | |||||||
| SwissProt | |||||||
| TrEMBL | |||||||
| UniProt Splice Variant | |||||||
| Entrez Gene | |||||||
| UniGene | Bt.84824 | ||||||
| RefSeq | NM_001193052 XM_005203240 XM_005203241 XM_005203242 | ||||||
| HUGO | |||||||
| OMIM | |||||||
| CCDS | |||||||
| HPRD | |||||||
| IMGT | |||||||
| EMBL | |||||||
| GenPept | |||||||
| RNA Seq Atlas | |||||||
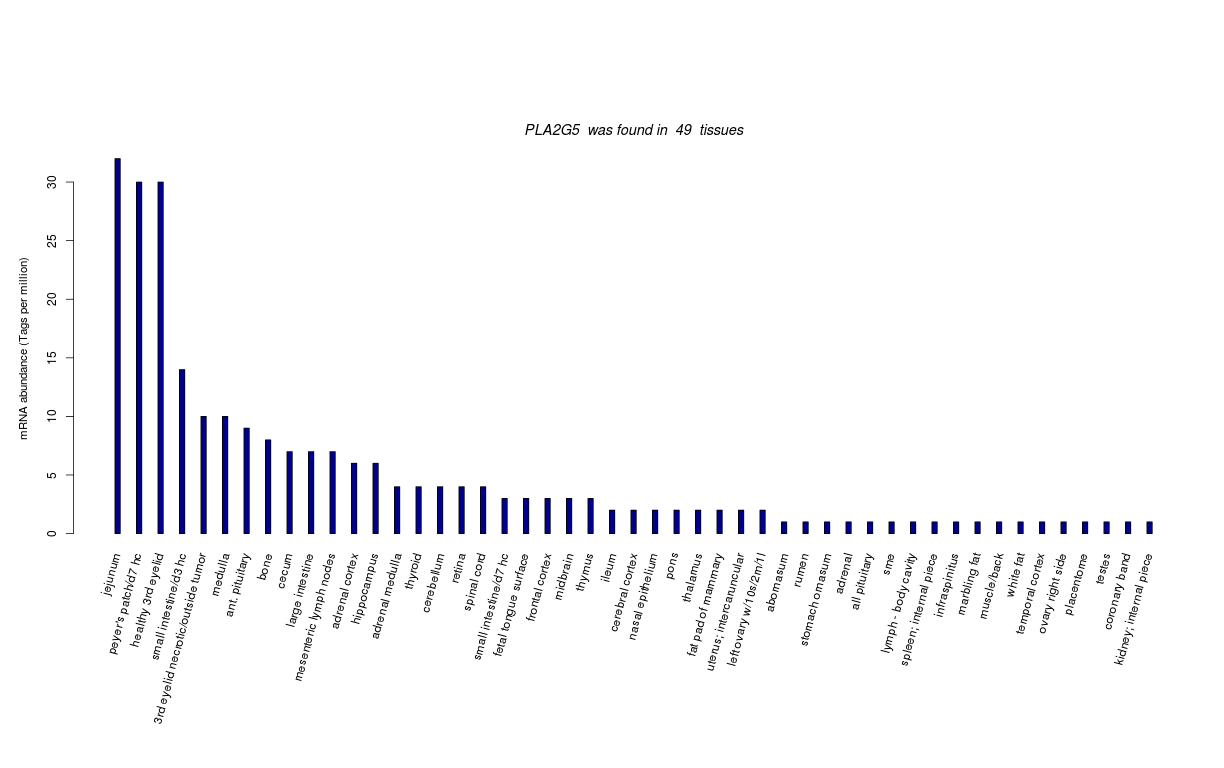
| Transcript Frequencies | |||||||
| Tag Count based mRNA-Abundances across 87 different Tissues (TPM).
Based on Data from Bovine Gene Atlas |
(Move your mouse over the image to view a more detailed version) |
||||||