| Bos taurus Gene: BT.30969 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Summary | |||||||||||
| InnateDB Gene | IDBG-647296.3 | ||||||||||
| Last Modified | 2014-10-13 [Report errors or provide feedback] | ||||||||||
| Gene Symbol | BT.30969 | ||||||||||
| Gene Name | UDP-GalNAc:beta-1,3-N-acetylgalactosaminyltransferase 1 | ||||||||||
| Synonyms | |||||||||||
| Species | Bos taurus | ||||||||||
| Ensembl Gene | ENSBTAG00000046104 | ||||||||||
| Encoded Proteins |
UDP-GalNAc:beta-1,3-N-acetylgalactosaminyltransferase 1
|
||||||||||
| Protein Structure | |||||||||||
| Useful resources | Stemformatics EHFPI ImmGen | ||||||||||
| Entrez Gene | |||||||||||
| Summary |
This gene does not have any Entrez summary - the following is the summary from its human ortholog ENSG00000169255:
This gene is a member of the beta-1,3-galactosyltransferase (beta3GalT) gene family. This family encodes type II membrane-bound glycoproteins with diverse enzymatic functions using different donor substrates (UDP-galactose and UDP-N-acetylglucosamine) and different acceptor sugars (N-acetylglucosamine, galactose, N-acetylgalactosamine). The beta3GalT genes are distantly related to the Drosophila Brainiac gene and have the protein coding sequence contained in a single exon. The beta3GalT proteins also contain conserved sequences not found in the beta4GalT or alpha3GalT proteins. The carbohydrate chains synthesized by these enzymes are designated as type 1, whereas beta4GalT enzymes synthesize type 2 carbohydrate chains. The ratio of type 1:type 2 chains changes during embryogenesis. By sequence similarity, the beta3GalT genes fall into at least two groups: beta3GalT4 and 4 other beta3GalT genes (beta3GalT1-3, beta3GalT5). The encoded protein of this gene does not use N-acetylglucosamine as an acceptor sugar at all. Multiple transcript variants that are alternatively spliced in the 5' UTR have been described; they all encode the same protein. [provided by RefSeq, Jul 2008] This gene is a member of the beta-1,3-galactosyltransferase (beta3GalT) gene family. This family encodes type II membrane-bound glycoproteins with diverse enzymatic functions using different donor substrates (UDP-galactose and UDP-N-acetylglucosamine) and different acceptor sugars (N-acetylglucosamine, galactose, N-acetylgalactosamine). The beta3GalT genes are distantly related to the Drosophila Brainiac gene and have the protein coding sequence contained in a single exon. The beta3GalT proteins also contain conserved sequences not found in the beta4GalT or alpha3GalT proteins. The carbohydrate chains synthesized by these enzymes are designated as type 1, whereas beta4GalT enzymes synthesize type 2 carbohydrate chains. The ratio of type 1:type 2 chains changes during embryogenesis. By sequence similarity, the beta3GalT genes fall into at least two groups: beta3GalT4 and 4 other beta3GalT genes (beta3GalT1-3, beta3GalT5). The encoded protein of this gene does not use N-acetylglucosamine as an acceptor sugar at all. Multiple transcript variants that are alternatively spliced in the 5\' UTR have been described; they all encode the same protein. [provided by RefSeq, Jul 2008] |
||||||||||
| Gene Information | |||||||||||
| Type | Protein coding | ||||||||||
| Genomic Location | Chromosome 1:107212961-107213956 | ||||||||||
| Strand | Forward strand | ||||||||||
| Band | |||||||||||
| Transcripts |
|
||||||||||
| Interactions | |||||||||||
| Number of Interactions |
This gene and/or its encoded proteins are associated with 0 experimentally validated interaction(s) in this database.
They are also associated with 1 interaction(s) predicted by orthology.
|
||||||||||
| Gene Ontology | |||||||||||
Molecular Function |
|
||||||||||
| Biological Process |
|
||||||||||
| Cellular Component |
|
||||||||||
| Orthologs | |||||||||||
|
Species
Homo sapiens
Mus musculus
|
Gene ID
Gene Order
|
||||||||||
| Pathway Predictions based on Human Orthology Data | |||||||||||
| NETPATH | |||||||||||
| REACTOME | |||||||||||
| KEGG |
Glycosphingolipid biosynthesis pathway
Glycosphingolipid biosynthesis pathway
|
||||||||||
| INOH | |||||||||||
| PID NCI | |||||||||||
| Cross-References | |||||||||||
| SwissProt | |||||||||||
| TrEMBL | |||||||||||
| UniProt Splice Variant | |||||||||||
| Entrez Gene | |||||||||||
| UniGene | Bt.30969 | ||||||||||
| RefSeq | NM_001076963 XM_005201746 XM_005201747 XM_005201748 XM_005201749 XM_005201750 XM_005201751 XM_005201752 XM_005201753 XM_005201754 XM_005201755 XM_005201756 XM_005201757 XM_005201758 XM_005201759 XM_005201760 XM_005201761 XM_005201762 | ||||||||||
| HUGO | |||||||||||
| OMIM | |||||||||||
| CCDS | |||||||||||
| HPRD | |||||||||||
| IMGT | |||||||||||
| EMBL | |||||||||||
| GenPept | |||||||||||
| RNA Seq Atlas | |||||||||||
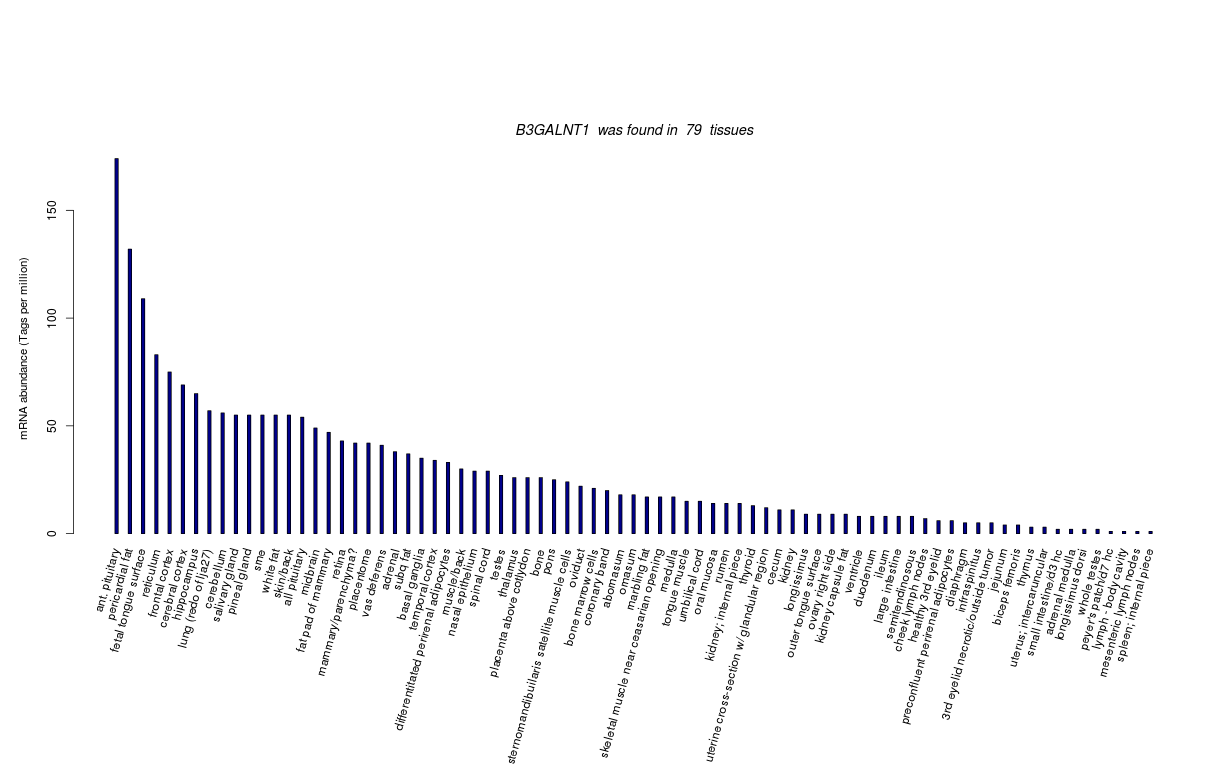
| Transcript Frequencies | |||||||||||
| Tag Count based mRNA-Abundances across 87 different Tissues (TPM).
Based on Data from Bovine Gene Atlas |
(Move your mouse over the image to view a more detailed version) |
||||||||||