| Bos taurus Gene: H1F0 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Summary | |||||||||||||||
| InnateDB Gene | IDBG-648421.3 | ||||||||||||||
| Last Modified | 2014-10-13 [Report errors or provide feedback] | ||||||||||||||
| Gene Symbol | H1F0 | ||||||||||||||
| Gene Name | Histone H1.0 | ||||||||||||||
| Synonyms | |||||||||||||||
| Species | Bos taurus | ||||||||||||||
| Ensembl Gene | ENSBTAG00000038439 | ||||||||||||||
| Encoded Proteins |
Histone H1.0
|
||||||||||||||
| Protein Structure | |||||||||||||||
| Useful resources | Stemformatics EHFPI ImmGen | ||||||||||||||
| Entrez Gene | |||||||||||||||
| Summary |
This gene does not have any Entrez summary - the following is the summary from its human ortholog ENSG00000189060:
Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Nucleosomes consist of approximately 146 bp of DNA wrapped around a histone octamer composed of pairs of each of the four core histones (H2A, H2B, H3, and H4). The chromatin fiber is further compacted through the interaction of a linker histone, H1, with the DNA between the nucleosomes to form higher order chromatin structures. This gene is intronless and encodes a member of the histone H1 family. [provided by RefSeq, Jul 2008] |
||||||||||||||
| Gene Information | |||||||||||||||
| Type | Protein coding | ||||||||||||||
| Genomic Location | Chromosome 5:110122673-110124878 | ||||||||||||||
| Strand | Forward strand | ||||||||||||||
| Band | |||||||||||||||
| Transcripts |
|
||||||||||||||
| Interactions | |||||||||||||||
| Number of Interactions |
This gene and/or its encoded proteins are associated with 0 experimentally validated interaction(s) in this database.
They are also associated with 20 interaction(s) predicted by orthology.
|
||||||||||||||
| Gene Ontology | |||||||||||||||
Molecular Function |
|
||||||||||||||
| Biological Process |
|
||||||||||||||
| Cellular Component |
|
||||||||||||||
| Orthologs | |||||||||||||||
|
Species
Homo sapiens
Mus musculus
|
Gene ID
Gene Order
|
||||||||||||||
| Pathways | |||||||||||||||
| NETPATH | |||||||||||||||
| REACTOME |
Apoptosis induced DNA fragmentation pathway
Cellular responses to stress pathway
Activation of DNA fragmentation factor pathway
Cellular Senescence pathway
Apoptotic execution phase pathway
Formation of Senescence-Associated Heterochromatin Foci (SAHF) pathway
DNA Damage/Telomere Stress Induced Senescence pathway
Apoptosis pathway
|
||||||||||||||
| KEGG | |||||||||||||||
| INOH | |||||||||||||||
| PID NCI | |||||||||||||||
| Pathway Predictions based on Human Orthology Data | |||||||||||||||
| NETPATH | |||||||||||||||
| REACTOME |
Activation of DNA fragmentation factor pathway
Apoptotic execution phase pathway
Cellular responses to stress pathway
Apoptosis induced DNA fragmentation pathway
Formation of Senescence-Associated Heterochromatin Foci (SAHF) pathway
Apoptosis pathway
DNA Damage/Telomere Stress Induced Senescence pathway
Cellular Senescence pathway
Apoptosis pathway
Cellular responses to stress pathway
Apoptosis induced DNA fragmentation pathway
DNA Damage/Telomere Stress Induced Senescence pathway
Activation of DNA fragmentation factor pathway
Cellular Senescence pathway
Formation of Senescence-Associated Heterochromatin Foci (SAHF) pathway
Apoptotic execution phase pathway
|
||||||||||||||
| KEGG | |||||||||||||||
| INOH | |||||||||||||||
| PID NCI | |||||||||||||||
| Cross-References | |||||||||||||||
| SwissProt | |||||||||||||||
| TrEMBL | |||||||||||||||
| UniProt Splice Variant | |||||||||||||||
| Entrez Gene | |||||||||||||||
| UniGene | Bt.74176 | ||||||||||||||
| RefSeq | NM_001076487 | ||||||||||||||
| HUGO | |||||||||||||||
| OMIM | |||||||||||||||
| CCDS | |||||||||||||||
| HPRD | |||||||||||||||
| IMGT | |||||||||||||||
| EMBL | |||||||||||||||
| GenPept | |||||||||||||||
| RNA Seq Atlas | |||||||||||||||
| Transcript Frequencies | |||||||||||||||
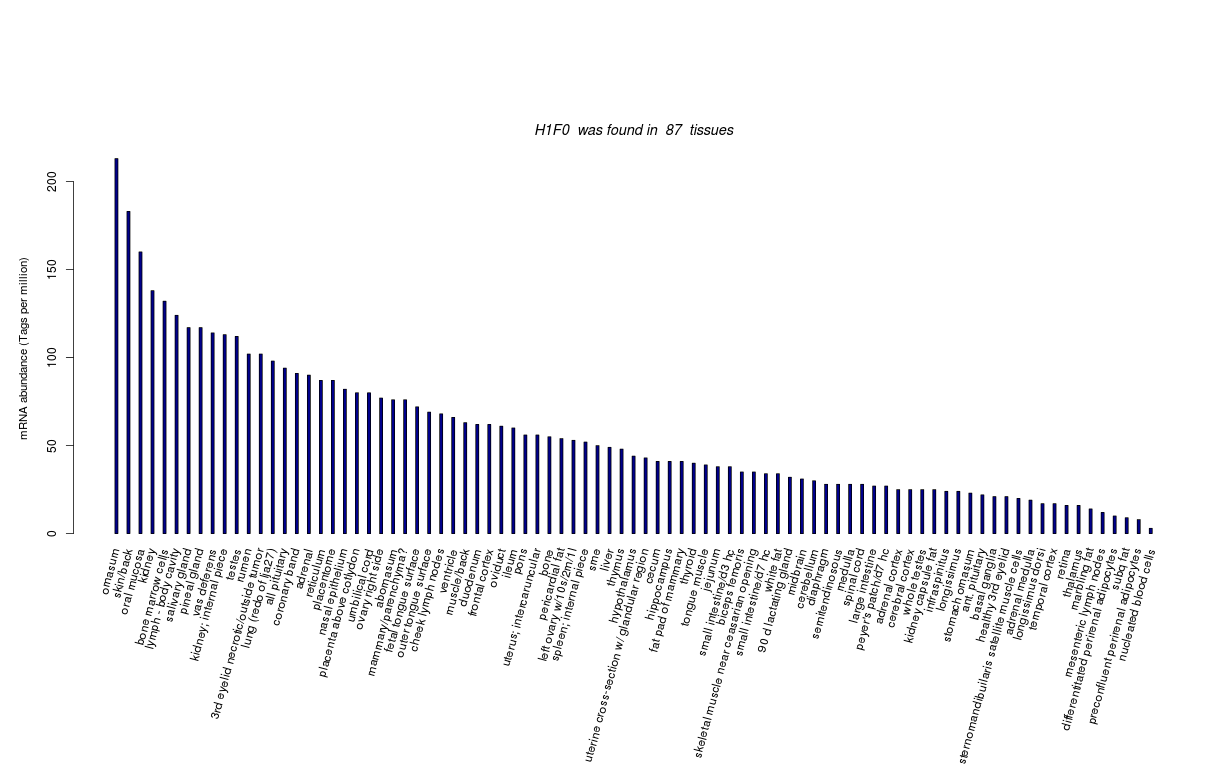
| Tag Count based mRNA-Abundances across 87 different Tissues (TPM).
Based on Data from Bovine Gene Atlas |
(Move your mouse over the image to view a more detailed version) |
||||||||||||||