| Bos taurus Gene: BT.45582 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Summary | |||||||||||
| InnateDB Gene | IDBG-642722.3 | ||||||||||
| Last Modified | 2014-10-13 [Report errors or provide feedback] | ||||||||||
| Gene Symbol | BT.45582 | ||||||||||
| Gene Name | chromobox protein homolog 2 | ||||||||||
| Synonyms | |||||||||||
| Species | Bos taurus | ||||||||||
| Ensembl Gene | ENSBTAG00000038306 | ||||||||||
| Encoded Proteins |
Uncharacterized protein
|
||||||||||
| Protein Structure | |||||||||||
| Useful resources | Stemformatics EHFPI ImmGen | ||||||||||
| Entrez Gene | |||||||||||
| Summary |
This gene does not have any Entrez summary - the following is the summary from its human ortholog ENSG00000173894:
This gene encodes a component of the polycomb multiprotein complex, which is required to maintain the transcriptionally repressive state of many genes throughout development via chromatin remodeling and modification of histones. Disruption of this gene in mice results in male-to-female gonadal sex reversal. Mutations in this gene are also associated with gonadal dysgenesis in humans. Alternatively spliced transcript variants encoding different isoforms have been noted for this gene.[provided by RefSeq, Mar 2010] |
||||||||||
| Gene Information | |||||||||||
| Type | Protein coding | ||||||||||
| Genomic Location | Chromosome 19:53371117-53377423 | ||||||||||
| Strand | Reverse strand | ||||||||||
| Band | |||||||||||
| Transcripts |
|
||||||||||
| Interactions | |||||||||||
| Number of Interactions |
This gene and/or its encoded proteins are associated with 0 experimentally validated interaction(s) in this database.
They are also associated with 43 interaction(s) predicted by orthology.
|
||||||||||
| Gene Ontology | |||||||||||
Molecular Function |
|
||||||||||
| Biological Process |
|
||||||||||
| Cellular Component |
|
||||||||||
| Orthologs | |||||||||||
|
Species
Homo sapiens
Mus musculus
|
Gene ID
Gene Order
|
||||||||||
| Pathway Predictions based on Human Orthology Data | |||||||||||
| NETPATH | |||||||||||
| REACTOME |
Cellular responses to stress pathway
Cellular Senescence pathway
Oxidative Stress Induced Senescence pathway
Oxidative Stress Induced Senescence pathway
Cellular responses to stress pathway
Cellular Senescence pathway
|
||||||||||
| KEGG | |||||||||||
| INOH | |||||||||||
| PID NCI | |||||||||||
| Cross-References | |||||||||||
| SwissProt | |||||||||||
| TrEMBL | |||||||||||
| UniProt Splice Variant | |||||||||||
| Entrez Gene | |||||||||||
| UniGene | Bt.45582 | ||||||||||
| RefSeq | NM_001205593 | ||||||||||
| HUGO | |||||||||||
| OMIM | |||||||||||
| CCDS | |||||||||||
| HPRD | |||||||||||
| IMGT | |||||||||||
| EMBL | |||||||||||
| GenPept | |||||||||||
| RNA Seq Atlas | |||||||||||
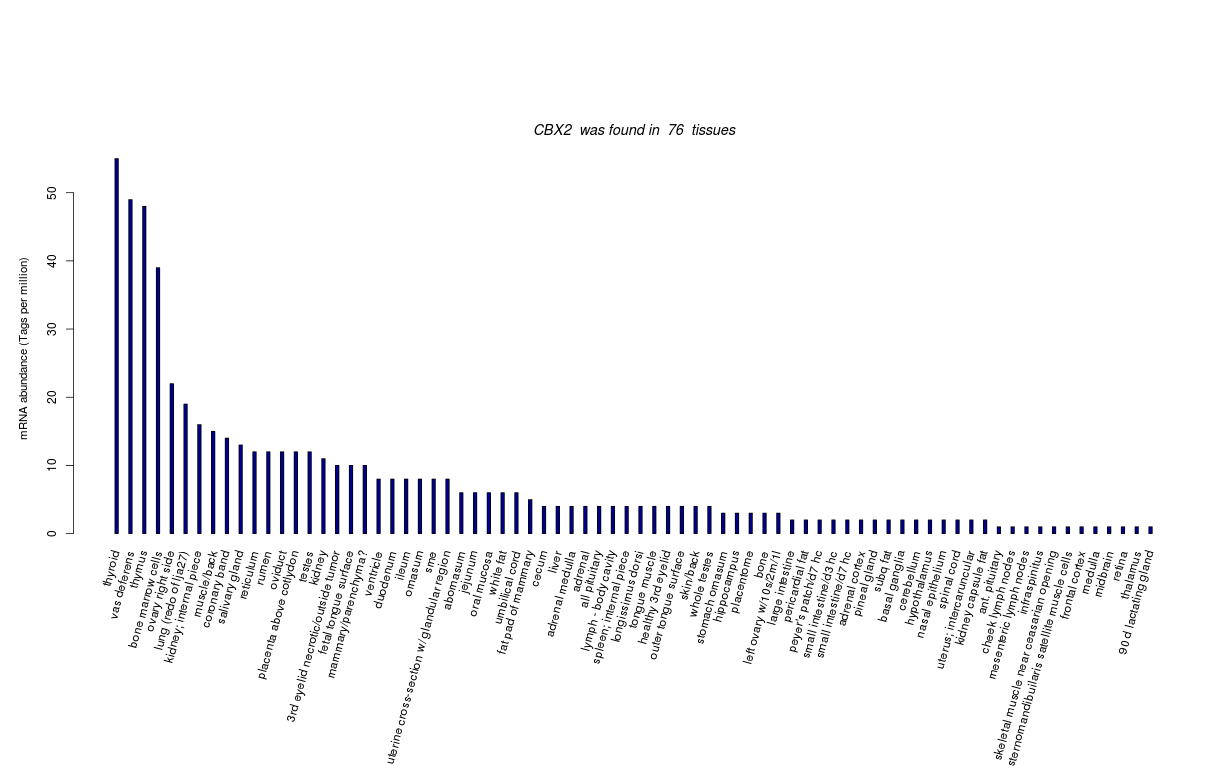
| Transcript Frequencies | |||||||||||
| Tag Count based mRNA-Abundances across 87 different Tissues (TPM).
Based on Data from Bovine Gene Atlas |
(Move your mouse over the image to view a more detailed version) |
||||||||||