| Bos taurus Gene: FPGT | |||||||
|---|---|---|---|---|---|---|---|
| Summary | |||||||
| InnateDB Gene | IDBG-637869.3 | ||||||
| Last Modified | 2014-10-13 [Report errors or provide feedback] | ||||||
| Gene Symbol | FPGT | ||||||
| Gene Name | fucose-1-phosphate guanylyltransferase | ||||||
| Synonyms | |||||||
| Species | Bos taurus | ||||||
| Ensembl Gene | ENSBTAG00000038480 | ||||||
| Encoded Proteins |
fucose-1-phosphate guanylyltransferase
|
||||||
| Protein Structure | |||||||
| Useful resources | Stemformatics EHFPI ImmGen | ||||||
| Entrez Gene | |||||||
| Summary |
This gene does not have any Entrez summary - the following is the summary from its human ortholog ENSG00000254685:
L-fucose is a key sugar in glycoproteins and other complex carbohydrates since it may be involved in many of the functional roles of these macromolecules, such as in cell-cell recognition. The fucosyl donor for these fucosylated oligosaccharides is GDP-beta-L-fucose. There are two alternate pathways for the biosynthesis of GDP-fucose; the major pathway converts GDP-alpha-D-mannose to GDP-beta-L-fucose. The protein encoded by this gene participates in an alternate pathway that is present in certain mammalian tissues, such as liver and kidney, and appears to function as a salvage pathway to reutilize L-fucose arising from the turnover of glycoproteins and glycolipids. This pathway involves the phosphorylation of L-fucose to form beta-L-fucose-1-phosphate, and then condensation of the beta-L-fucose-1-phosphate with GTP by fucose-1-phosphate guanylyltransferase to form GDP-beta-L-fucose. Alternative splicing results in multiple transcript variants. Read-through transcription also exists between this gene and the neighboring downstream TNNI3 interacting kinase (TNNI3K) gene. [provided by RefSeq, Dec 2010] |
||||||
| Gene Information | |||||||
| Type | Protein coding | ||||||
| Genomic Location | Chromosome 3:70878977-70887093 | ||||||
| Strand | Reverse strand | ||||||
| Band | |||||||
| Transcripts |
|
||||||
| Interactions | |||||||
| Number of Interactions |
This gene and/or its encoded proteins are associated with 0 experimentally validated interaction(s) in this database.
They are also associated with 2 interaction(s) predicted by orthology.
|
||||||
| Gene Ontology | |||||||
Molecular Function |
|
||||||
| Biological Process |
|
||||||
| Cellular Component |
|
||||||
| Orthologs | |||||||
|
Species
Mus musculus
Homo sapiens
|
Gene ID
Gene Order
|
||||||
| Pathway Predictions based on Human Orthology Data | |||||||
| NETPATH | |||||||
| REACTOME | |||||||
| KEGG |
Amino sugar and nucleotide sugar metabolism pathway
Fructose and mannose metabolism pathway
Amino sugar and nucleotide sugar metabolism pathway
Fructose and mannose metabolism pathway
|
||||||
| INOH |
Fructose Mannose metabolism pathway
|
||||||
| PID NCI | |||||||
| Cross-References | |||||||
| SwissProt | |||||||
| TrEMBL | E1B9U5 | ||||||
| UniProt Splice Variant | |||||||
| Entrez Gene | 100138313 | ||||||
| UniGene | Bt.107926 Bt.17095 Bt.67968 | ||||||
| RefSeq | NM_001191283 | ||||||
| HUGO | HGNC:3825 | ||||||
| OMIM | |||||||
| CCDS | |||||||
| HPRD | |||||||
| IMGT | |||||||
| EMBL | DAAA02008315 | ||||||
| GenPept | |||||||
| RNA Seq Atlas | 100138313 | ||||||
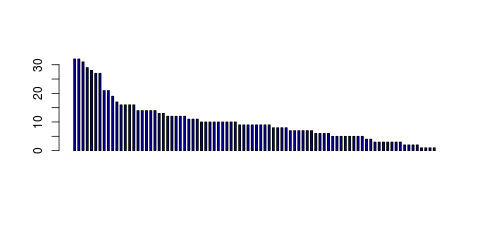
| Transcript Frequencies | |||||||
| Tag Count based mRNA-Abundances across 87 different Tissues (TPM).
Based on Data from Bovine Gene Atlas |
(Move your mouse over the image to view a more detailed version) |
||||||