| Bos taurus Gene: BT.96672 | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Summary | |||||||||||||||||||||||||
| InnateDB Gene | IDBG-645560.3 | ||||||||||||||||||||||||
| Last Modified | 2014-10-13 [Report errors or provide feedback] | ||||||||||||||||||||||||
| Gene Symbol | BT.96672 | ||||||||||||||||||||||||
| Gene Name | interferon regulatory factor 7 | ||||||||||||||||||||||||
| Synonyms | |||||||||||||||||||||||||
| Species | Bos taurus | ||||||||||||||||||||||||
| Ensembl Gene | ENSBTAG00000047680 | ||||||||||||||||||||||||
| Encoded Proteins |
interferon regulatory factor 7
|
||||||||||||||||||||||||
| Protein Structure | |||||||||||||||||||||||||
| Useful resources | Stemformatics EHFPI ImmGen | ||||||||||||||||||||||||
| InnateDB Annotation from Orthologs | |||||||||||||||||||||||||
| Summary |
[Homo sapiens] IRF7 increases the expression of a broad range of IFN-stimulated genes including immunomodulatory cytokines and genes involved in antigen processing and presentation.
[Homo sapiens] IRF7 is activated in response to virus infection and stimulates the transcription of a set of cellular genes involved in host antiviral defence.
[Homo sapiens] IRF7 forms a complex with MYD88 and TRAF6 and this complex formation, as well as TRAF6-dependent IRF7 ubiquitination, is required for TLR-mediated interferon (IFN)-alpha induction.
[Homo sapiens] IRF7 is involved in the innate immune recognition of Plasmodium falciparum AT-rich DNA and in the subsequent induction of type I IFNs. Mice lacking Irf3/Irf7 are resistant to otherwise lethal cerebral malaria. (Demonstrated in mouse)
[Homo sapiens] Coronavirus engages papain-like proteases to escape from the innate antiviral response of the host by inhibiting TP53-IRF7-IFNB1 signalling.
[Homo sapiens] Paramyxoviruses trigger the DNA-damage response, a pathway required for RPS6KA5 activation of phospho Ser 276 RELA formation to trigger the IRF7-DDX58 amplification loop necessary for mucosal interferon production.
[Homo sapiens] AIP is a novel inhibitor of IRF7 and a negative regulator of innate antiviral signalling.
[Mus musculus] Irf7 is involved in the innate immune recognition of Plasmodium falciparum AT-rich DNA and in the subsequent induction of type I IFNs. Mice lacking Irf3/Irf7 are resistant to otherwise lethal cerebral malaria.
[Mus musculus] Plasmodium RNA is a pathogen-associated molecular pattern (PAMP) capable of activating a type I IFN response via the cytosolic pattern recognition receptors Ifih1 and Mavs, as well as via transcription factors Irf3 and Irf7.
[Mus musculus] Genetic deletion of Eif4ebp1 or Eif4ebp2 potentiates innate antiviral immunity by enhancing translation of Irf7.
[Mus musculus] DNA vaccine-induced, Irf7-dependent signalling, as part of the Tmem173 (Sting) pathway, is critical for generation of both innate cytokine signalling and antigen-specific B and T cell responses.
|
||||||||||||||||||||||||
| Entrez Gene | |||||||||||||||||||||||||
| Summary |
This gene does not have any Entrez summary - the following is the summary from its human ortholog ENSG00000185507:
IRF7 encodes interferon regulatory factor 7, a member of the interferon regulatory transcription factor (IRF) family. IRF7 has been shown to play a role in the transcriptional activation of virus-inducible cellular genes, including interferon beta chain genes. Inducible expression of IRF7 is largely restricted to lymphoid tissue. Multiple IRF7 transcript variants have been identified, although the functional consequences of these have not yet been established. [provided by RefSeq, Jul 2008] |
||||||||||||||||||||||||
| Gene Information | |||||||||||||||||||||||||
| Type | Protein coding | ||||||||||||||||||||||||
| Genomic Location | Chromosome 29:50903719-50906820 | ||||||||||||||||||||||||
| Strand | Forward strand | ||||||||||||||||||||||||
| Band | |||||||||||||||||||||||||
| Transcripts |
|
||||||||||||||||||||||||
| Interactions | |||||||||||||||||||||||||
| Number of Interactions |
This gene and/or its encoded proteins are associated with 0 experimentally validated interaction(s) in this database.
They are also associated with 63 interaction(s) predicted by orthology.
|
||||||||||||||||||||||||
| Gene Ontology | |||||||||||||||||||||||||
Molecular Function |
|
||||||||||||||||||||||||
| Biological Process |
|
||||||||||||||||||||||||
| Cellular Component |
|
||||||||||||||||||||||||
| Orthologs | |||||||||||||||||||||||||
|
Species
Homo sapiens
Mus musculus
|
Gene ID
Gene Order
|
||||||||||||||||||||||||
| Pathway Predictions based on Human Orthology Data | |||||||||||||||||||||||||
| NETPATH | |||||||||||||||||||||||||
| REACTOME |
TRAF6 mediated IRF7 activation pathway
TRAF3-dependent IRF activation pathway pathway
RIG-I/MDA5 mediated induction of IFN-alpha/beta pathways pathway
Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon pathway
MyD88-independent cascade pathway
Toll Like Receptor 3 (TLR3) Cascade pathway
TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling pathway
MyD88 dependent cascade initiated on endosome pathway
Toll Like Receptor 9 (TLR9) Cascade pathway
Toll Like Receptor 4 (TLR4) Cascade pathway
Interferon alpha/beta signaling pathway
Interferon gamma signaling pathway
Factors involved in megakaryocyte development and platelet production pathway
Toll Like Receptor 7/8 (TLR7/8) Cascade pathway
Cytokine Signaling in Immune system pathway
Innate Immune System pathway
Toll-Like Receptors Cascades pathway
Interferon Signaling pathway
DEx/H-box helicases activate type I IFN and inflammatory cytokines production pathway
Immune System pathway
Activated TLR4 signalling pathway
TRIF-mediated TLR3/TLR4 signaling pathway
Cytosolic sensors of pathogen-associated DNA pathway
Hemostasis pathway
Toll Like Receptor 3 (TLR3) Cascade pathway
Innate Immune System pathway
Hemostasis pathway
Cytokine Signaling in Immune system pathway
Immune System pathway
TRAF6 mediated IRF7 activation pathway
TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling pathway
Factors involved in megakaryocyte development and platelet production pathway
Toll Like Receptor 9 (TLR9) Cascade pathway
Interferon gamma signaling pathway
Interferon Signaling pathway
DEx/H-box helicases activate type I IFN and inflammatory cytokines production pathway
Toll-Like Receptors Cascades pathway
MyD88 dependent cascade initiated on endosome pathway
Cytosolic sensors of pathogen-associated DNA pathway
Activated TLR4 signalling pathway
MyD88-independent cascade pathway
RIG-I/MDA5 mediated induction of IFN-alpha/beta pathways pathway
Toll Like Receptor 7/8 (TLR7/8) Cascade pathway
TRIF-mediated TLR3/TLR4 signaling pathway
TRAF3-dependent IRF activation pathway pathway
Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon pathway
Toll Like Receptor 4 (TLR4) Cascade pathway
|
||||||||||||||||||||||||
| KEGG |
Toll-like receptor signaling pathway pathway
RIG-I-like receptor signaling pathway pathway
Cytosolic DNA-sensing pathway pathway
Hepatitis C pathway
Toll-like receptor signaling pathway pathway
RIG-I-like receptor signaling pathway pathway
Cytosolic DNA-sensing pathway pathway
Hepatitis C pathway
|
||||||||||||||||||||||||
| INOH | |||||||||||||||||||||||||
| PID NCI |
Regulation of nuclear SMAD2/3 signaling
|
||||||||||||||||||||||||
| Cross-References | |||||||||||||||||||||||||
| SwissProt | |||||||||||||||||||||||||
| TrEMBL | G3N3Q5 | ||||||||||||||||||||||||
| UniProt Splice Variant | |||||||||||||||||||||||||
| Entrez Gene | 100125591 | ||||||||||||||||||||||||
| UniGene | Bt.96672 | ||||||||||||||||||||||||
| RefSeq | NM_001105040 XM_005227253 XM_005227254 | ||||||||||||||||||||||||
| HUGO | HGNC:6122 | ||||||||||||||||||||||||
| OMIM | |||||||||||||||||||||||||
| CCDS | |||||||||||||||||||||||||
| HPRD | |||||||||||||||||||||||||
| IMGT | |||||||||||||||||||||||||
| EMBL | DAAA02063766 | ||||||||||||||||||||||||
| GenPept | |||||||||||||||||||||||||
| RNA Seq Atlas | 100125591 | ||||||||||||||||||||||||
| Transcript Frequencies | |||||||||||||||||||||||||
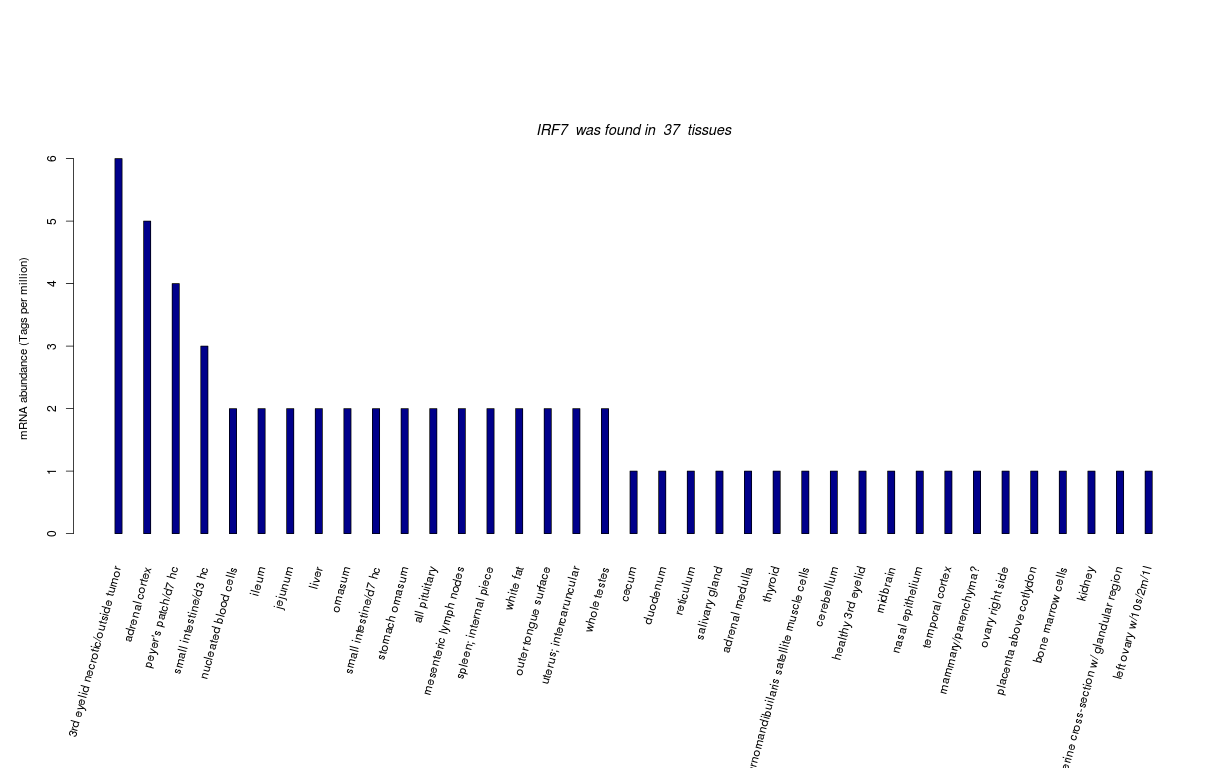
| Tag Count based mRNA-Abundances across 87 different Tissues (TPM).
Based on Data from Bovine Gene Atlas |
(Move your mouse over the image to view a more detailed version) |
||||||||||||||||||||||||