| Bos taurus Gene: P2RY2 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Summary | |||||||||||||
| InnateDB Gene | IDBG-635267.3 | ||||||||||||
| Last Modified | 2014-10-13 [Report errors or provide feedback] | ||||||||||||
| Gene Symbol | P2RY2 | ||||||||||||
| Gene Name | P2Y purinoceptor 2 | ||||||||||||
| Synonyms | |||||||||||||
| Species | Bos taurus | ||||||||||||
| Ensembl Gene | ENSBTAG00000039050 | ||||||||||||
| Encoded Proteins |
P2Y purinoceptor 2
|
||||||||||||
| Protein Structure | |||||||||||||
| Useful resources | Stemformatics EHFPI ImmGen | ||||||||||||
| Entrez Gene | |||||||||||||
| Summary |
This gene does not have any Entrez summary - the following is the summary from its human ortholog ENSG00000175591:
The product of this gene belongs to the family of G-protein coupled receptors. This family has several receptor subtypes with different pharmacological selectivity, which overlaps in some cases, for various adenosine and uridine nucleotides. This receptor is responsive to both adenosine and uridine nucleotides. It may participate in control of the cell cycle of endometrial carcinoma cells. Three transcript variants encoding the same protein have been identified for this gene. [provided by RefSeq, Jul 2008] The product of this gene belongs to the family of P2 receptors, which is activated by extracellular nucleotides and subdivided into P2X ligand-gated ion channels and P2Y G-protein coupled receptors. This family has several receptor subtypes with different pharmacological selectivity, which overlaps in some cases, for various adenosine and uridine nucleotides. This receptor, found on many cell types, is activated by ATP and UTP and is reported to be overexpressed on some cancer cell types. It is involved in many cellular functions, such as proliferation, apoptosis and inflammation. Three transcript variants encoding the same protein have been identified for this gene. [provided by RefSeq, Mar 2013] |
||||||||||||
| Gene Information | |||||||||||||
| Type | Protein coding | ||||||||||||
| Genomic Location | Chromosome 15:53492850-53499415 | ||||||||||||
| Strand | Forward strand | ||||||||||||
| Band | |||||||||||||
| Transcripts |
|
||||||||||||
| Interactions | |||||||||||||
| Number of Interactions |
This gene and/or its encoded proteins are associated with 0 experimentally validated interaction(s) in this database.
They are also associated with 2 interaction(s) predicted by orthology.
|
||||||||||||
| Gene Ontology | |||||||||||||
Molecular Function |
|
||||||||||||
| Biological Process |
|
||||||||||||
| Cellular Component |
|
||||||||||||
| Orthologs | |||||||||||||
|
Species
Homo sapiens
Mus musculus
|
Gene ID
Gene Order
|
||||||||||||
| Pathway Predictions based on Human Orthology Data | |||||||||||||
| NETPATH | |||||||||||||
| REACTOME |
G alpha (q) signalling events pathway
Gastrin-CREB signalling pathway via PKC and MAPK pathway
P2Y receptors pathway
Class A/1 (Rhodopsin-like receptors) pathway
Signaling by GPCR pathway
Signal Transduction pathway
GPCR downstream signaling pathway
GPCR ligand binding pathway
Nucleotide-like (purinergic) receptors pathway
P2Y receptors pathway
GPCR downstream signaling pathway
Signaling by GPCR pathway
Class A/1 (Rhodopsin-like receptors) pathway
Gastrin-CREB signalling pathway via PKC and MAPK pathway
Nucleotide-like (purinergic) receptors pathway
GPCR ligand binding pathway
G alpha (q) signalling events pathway
Signal Transduction pathway
|
||||||||||||
| KEGG |
Neuroactive ligand-receptor interaction pathway
Neuroactive ligand-receptor interaction pathway
|
||||||||||||
| INOH | |||||||||||||
| PID NCI | |||||||||||||
| Cross-References | |||||||||||||
| SwissProt | |||||||||||||
| TrEMBL | |||||||||||||
| UniProt Splice Variant | |||||||||||||
| Entrez Gene | |||||||||||||
| UniGene | Bt.13024 | ||||||||||||
| RefSeq | NM_001166525 XM_005216164 | ||||||||||||
| HUGO | |||||||||||||
| OMIM | |||||||||||||
| CCDS | |||||||||||||
| HPRD | |||||||||||||
| IMGT | |||||||||||||
| EMBL | |||||||||||||
| GenPept | |||||||||||||
| RNA Seq Atlas | |||||||||||||
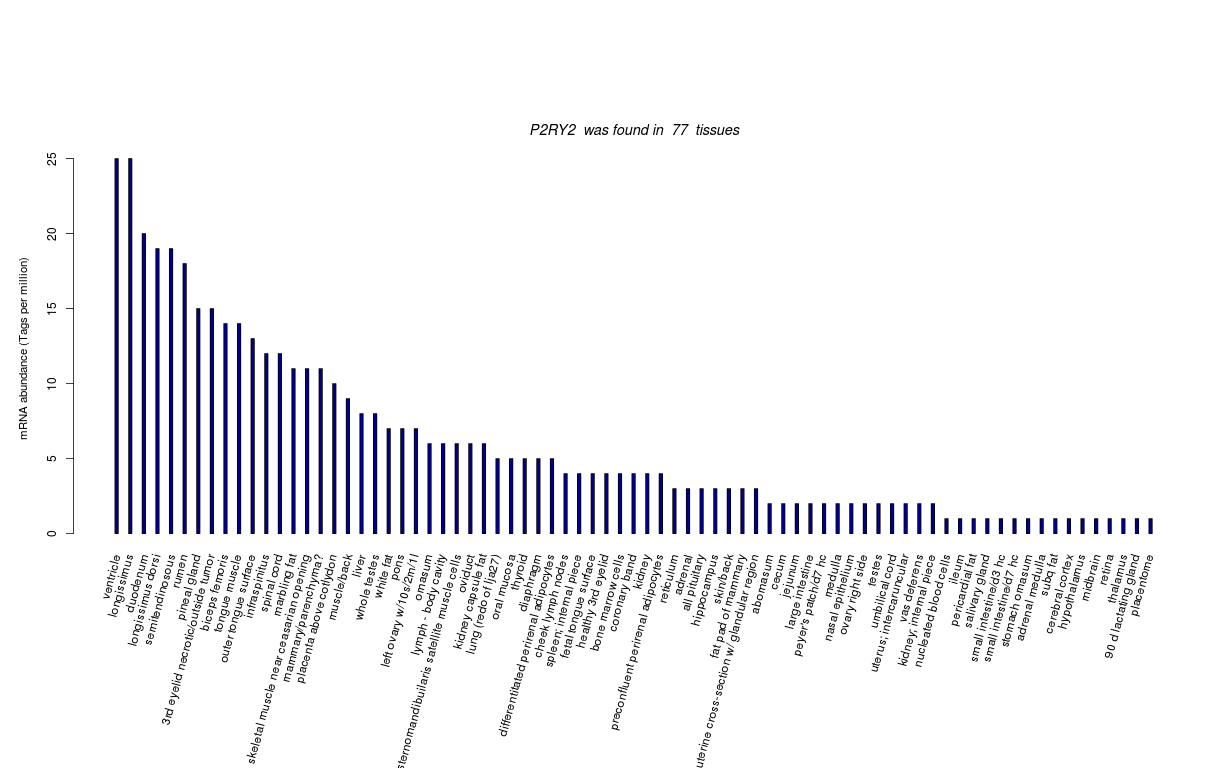
| Transcript Frequencies | |||||||||||||
| Tag Count based mRNA-Abundances across 87 different Tissues (TPM).
Based on Data from Bovine Gene Atlas |
(Move your mouse over the image to view a more detailed version) |
||||||||||||