| Bos taurus Gene: BT.35830 | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Summary | |||||||||||||||||||
| InnateDB Gene | IDBG-639593.3 | ||||||||||||||||||
| Last Modified | 2014-10-13 [Report errors or provide feedback] | ||||||||||||||||||
| Gene Symbol | BT.35830 | ||||||||||||||||||
| Gene Name | integrin alpha-M precursor | ||||||||||||||||||
| Synonyms | CD11B | ||||||||||||||||||
| Species | Bos taurus | ||||||||||||||||||
| Ensembl Gene | ENSBTAG00000047238 | ||||||||||||||||||
| Encoded Proteins |
integrin alpha M
|
||||||||||||||||||
| Protein Structure | |||||||||||||||||||
| Useful resources | Stemformatics EHFPI ImmGen | ||||||||||||||||||
| InnateDB Annotation from Orthologs | |||||||||||||||||||
| Summary |
[Homo sapiens] ITGAM (CD11b integrin) is activated via Toll-like receptors (TLRs) and engages in crosstalk with the MYD88 and TICAM1 (TRIF) pathways inhibiting TLR signalling in innate immune responses.
[Homo sapiens] ITGAM :: ITGB2 is the principal leukocyte receptor involved in the recognition of the fungus Candida albicans. Recognition of Pra1p protein of C. albicans by ITGAM :: ITGB2 plays a pivotal role in determining fungal virulence, and host response/protection against C. albicans infection. (Demonstrated in murine model)
[Mus musculus] Itgam (Cd11b integrin) is activated via Toll-like receptors (TLRs) and engages in crosstalk with the Myd88 and Ticam1 (TRIF) pathways inhibiting TLR signalling in innate immune responses.
[Mus musculus] Itgam :: Itgb2 is the principal leukocyte receptor involved in the recognition of the fungus Candida albicans. Recognition of Pra1p protein of C. albicans by Itgam :: Itgb2 plays a pivotal role in determining fungal virulence, and host response/protection against C. albicans infection.
[Mus musculus] Itgam (Cd11b) fine tunes the balance between adaptive and innate immune responses initiated by LPS by modulating the trafficking and signalling functions of Tlr4 in a cell-type-specific manner.
[Mus musculus] Ptpn11 phosphatase function positively regulates Clec7a- and Itgam-stimulated reactive oxygen species production in macrophages by dephosphorylating and thus mitigating the inhibitory function of Sirpa and by promoting Mapk1/Mapk3 activation.
|
||||||||||||||||||
| Entrez Gene | |||||||||||||||||||
| Summary |
This gene does not have any Entrez summary - the following is the summary from its human ortholog ENSG00000169896:
This gene encodes the integrin alpha M chain. Integrins are heterodimeric integral membrane proteins composed of an alpha chain and a beta chain. This I-domain containing alpha integrin combines with the beta 2 chain (ITGB2) to form a leukocyte-specific integrin referred to as macrophage receptor 1 ('Mac-1'), or inactivated-C3b (iC3b) receptor 3 ('CR3'). The alpha M beta 2 integrin is important in the adherence of neutrophils and monocytes to stimulated endothelium, and also in the phagocytosis of complement coated particles. Multiple transcript variants encoding different isoforms have been found for this gene. [provided by RefSeq, Mar 2009] This gene encodes the integrin alpha M chain. Integrins are heterodimeric integral membrane proteins composed of an alpha chain and a beta chain. This I-domain containing alpha integrin combines with the beta 2 chain (ITGB2) to form a leukocyte-specific integrin referred to as macrophage receptor 1 (\'Mac-1\'), or inactivated-C3b (iC3b) receptor 3 (\'CR3\'). The alpha M beta 2 integrin is important in the adherence of neutrophils and monocytes to stimulated endothelium, and also in the phagocytosis of complement coated particles. Multiple transcript variants encoding different isoforms have been found for this gene. [provided by RefSeq, Mar 2009] |
||||||||||||||||||
| Gene Information | |||||||||||||||||||
| Type | Protein coding | ||||||||||||||||||
| Genomic Location | Chromosome 25:27602222-27639821 | ||||||||||||||||||
| Strand | Forward strand | ||||||||||||||||||
| Band | |||||||||||||||||||
| Transcripts |
|
||||||||||||||||||
| Interactions | |||||||||||||||||||
| Number of Interactions |
This gene and/or its encoded proteins are associated with 0 experimentally validated interaction(s) in this database.
They are also associated with 15 interaction(s) predicted by orthology.
|
||||||||||||||||||
| Gene Ontology | |||||||||||||||||||
Molecular Function |
|
||||||||||||||||||
| Biological Process |
|
||||||||||||||||||
| Cellular Component |
|
||||||||||||||||||
| Orthologs | |||||||||||||||||||
|
Species
Homo sapiens
Mus musculus
|
Gene ID
Gene Order
|
||||||||||||||||||
| Pathway Predictions based on Human Orthology Data | |||||||||||||||||||
| NETPATH | |||||||||||||||||||
| REACTOME |
Toll Like Receptor 4 (TLR4) Cascade pathway
Cell surface interactions at the vascular wall pathway
Integrin cell surface interactions pathway
Extracellular matrix organization pathway
Innate Immune System pathway
Toll-Like Receptors Cascades pathway
Immune System pathway
Hemostasis pathway
Innate Immune System pathway
Hemostasis pathway
Cell surface interactions at the vascular wall pathway
Immune System pathway
Extracellular matrix organization pathway
Toll-Like Receptors Cascades pathway
Integrin cell surface interactions pathway
Toll Like Receptor 4 (TLR4) Cascade pathway
|
||||||||||||||||||
| KEGG |
Regulation of actin cytoskeleton pathway
Hematopoietic cell lineage pathway
Cell adhesion molecules (CAMs) pathway
Leukocyte transendothelial migration pathway
Leishmaniasis pathway
Staphylococcus aureus infection pathway
Amoebiasis pathway
Phagosome pathway
Cell adhesion molecules (CAMs) pathway
Hematopoietic cell lineage pathway
Regulation of actin cytoskeleton pathway
Leukocyte transendothelial migration pathway
Staphylococcus aureus infection pathway
Leishmaniasis pathway
Amoebiasis pathway
Phagosome pathway
|
||||||||||||||||||
| INOH |
Integrin signaling pathway pathway
Integrin signaling pathway pathway
|
||||||||||||||||||
| PID NCI |
Integrin family cell surface interactions
Beta2 integrin cell surface interactions
Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling
amb2 Integrin signaling
|
||||||||||||||||||
| Cross-References | |||||||||||||||||||
| SwissProt | |||||||||||||||||||
| TrEMBL | G3MYD9 | ||||||||||||||||||
| UniProt Splice Variant | |||||||||||||||||||
| Entrez Gene | 407124 | ||||||||||||||||||
| UniGene | Bt.35830 | ||||||||||||||||||
| RefSeq | NM_001039957 | ||||||||||||||||||
| HUGO | HGNC:6149 | ||||||||||||||||||
| OMIM | |||||||||||||||||||
| CCDS | |||||||||||||||||||
| HPRD | |||||||||||||||||||
| IMGT | |||||||||||||||||||
| EMBL | DAAA02057921 | ||||||||||||||||||
| GenPept | |||||||||||||||||||
| RNA Seq Atlas | 407124 | ||||||||||||||||||
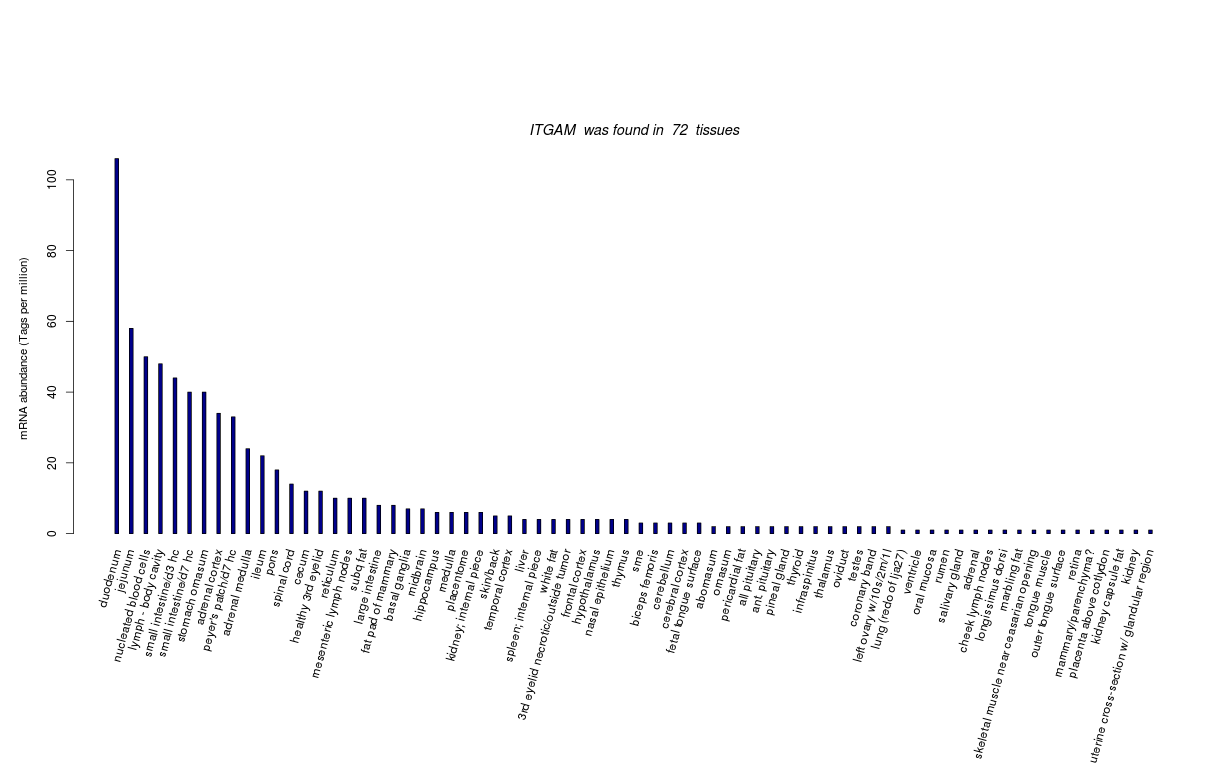
| Transcript Frequencies | |||||||||||||||||||
| Tag Count based mRNA-Abundances across 87 different Tissues (TPM).
Based on Data from Bovine Gene Atlas |
(Move your mouse over the image to view a more detailed version) |
||||||||||||||||||